TALLERS
Assignment for Systems Biology course; molecular level; University of Iceland 02/2023
How a drug for COVID-19 is designed?
- Introduction to the assignment
- Briefing on the mechanism of action of SARS-CoV-2
- Assignment questions
- References
Introduction to the assignment
In this assignment for the LVF601M course on Systems Biology at the University of Iceland, we will get some hints on the rational design of a new drug. In particular, we will visualize some simple details on the way an antiviral for SARS-CoV-2 can be designed.
Needed software and databases:
- OpenBabel to convert small molecules coordinates from one format to another. You can also use its online version.
- Chimera to visualize proteins and their interactions.
- UNIPROT to obtain information about a particular protein (sequence, structure, interactions…).
- PDB to obtain 3D structures of proteins.
Briefing on the mechanism of action of SARS-CoV-2
The SARS-CoV-2 virus is the cause of the disease known as COVID19. The Spike protein is responsible for anchoring the virus to the cell surface.
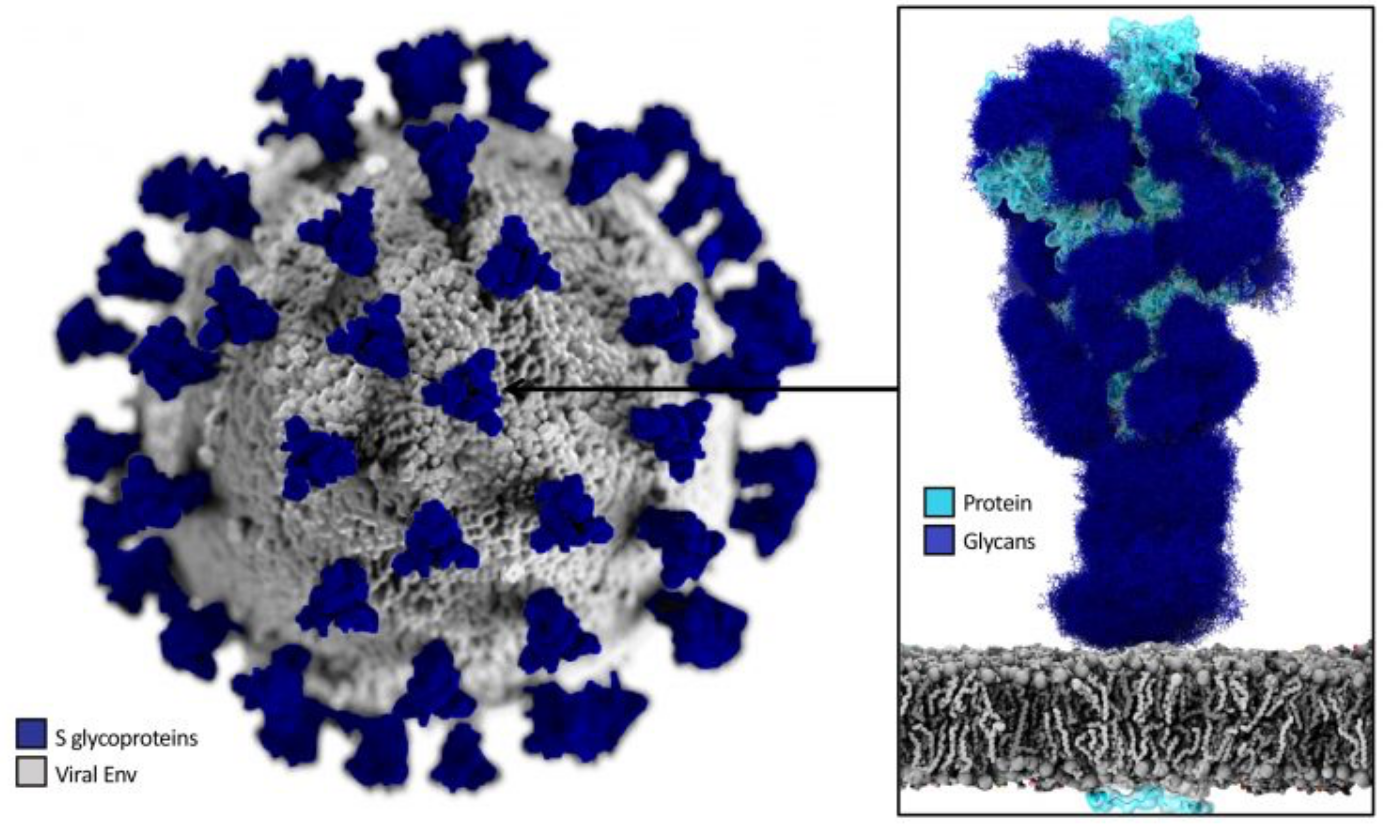 |
|---|
| Detail of the structure of the SARS-CoV-2, showing the Spike proteins in their glycosilated form |
From here the fusion of the membranes occurs and the virus pours its RNA content into the cell. This RNA uses the cellular machinery to replicate the virus and to generate many more that can infect other cells.
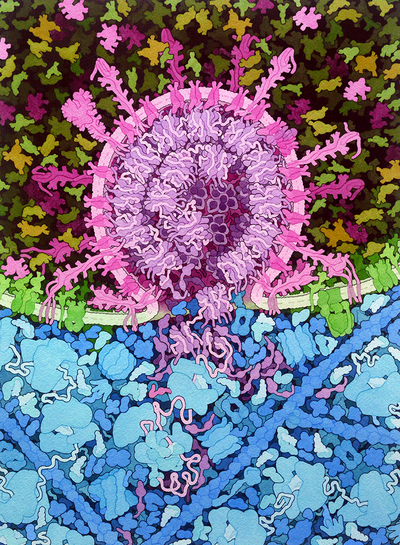 |
|---|
| Credit: David S. Goodsell |
A nice introductory video on the wat SARS-CoV-2 infects a cell and on the development of vaccines against it:
The virus machinery includes the information to translate a collection of proteins that, once assembled into new viral particles to infect new host cells. Some of these proteins are sructural and other have a precise funcion to help the maturation of the virus within the cell. See the details of the SARS-CoC-2 proteome in the figure.
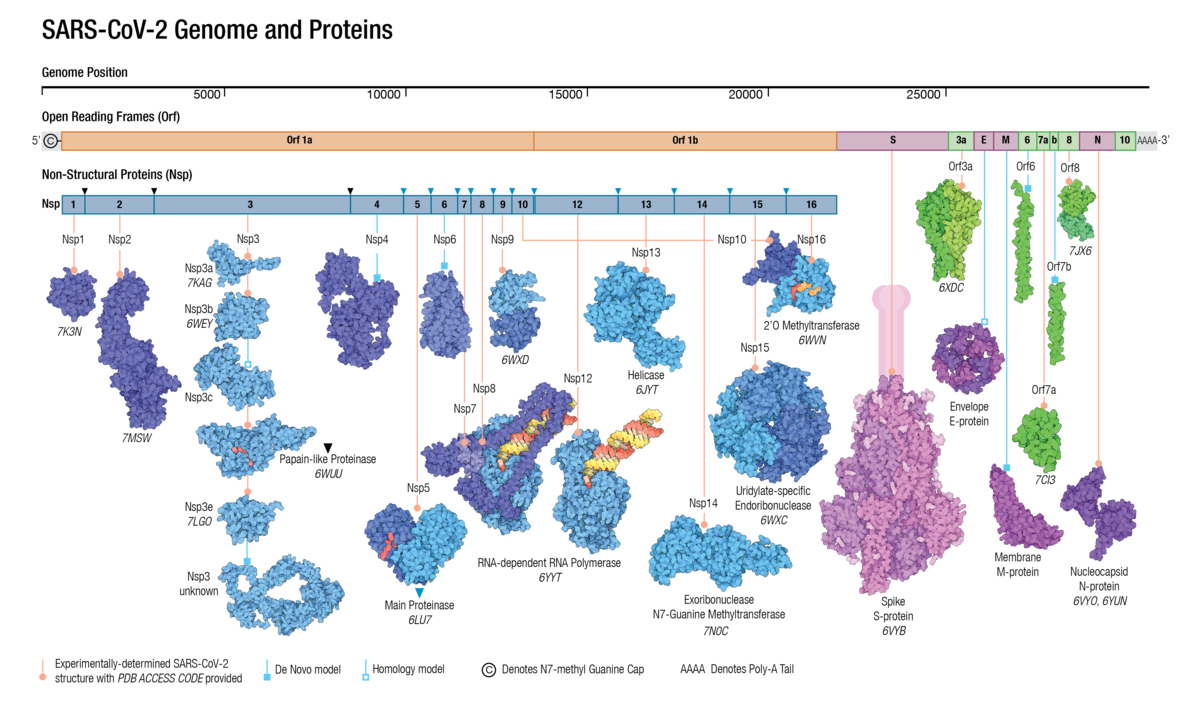 |
|---|
| The proteome of SARS-CoV-2 |
Assignment questions
1) Virus-cell interactions
- Get the human SARS-CoV-2 virus Spike protein Uniprot code.
- Identify the name of the cell surface protein that the SARS-CoV-2 Spike protein interacts with.
- Look in the PDB database for a structure of the complex between Spike protein and the above cell membrane protein. Add an image of the PDB obtained with Chimera.
- Look in the PDB database for a structure of the complex between Spike protein and antibodies. Add an image of the PDB obtained with Chimera.
- Identify the residues that are in the interface regions, using the
select zonetool in Chimera. Are they many? What do you think a good strategy for preventing SARS-CoV-2 to interact with the cell could be? Are the regions of interaction the same in the complexes you located in the above steps?
2) Variability in the SARS-CoV-2 genome
Go to the SARS-CoV-2 genome variation site at Stanford University: COVDB. Look for the page devoted to the omicron variants:
- Is the variability homogeneous? why do you think it is like this in terms of viral-host interaction evolution?
- Check in particular the genomic region for 3CLpro. Can you give a rough measure of the percentage of variation of Spike and 3CLpro?
3) Rational Drug Discovery
Ligand
- Go to the DRUGBANK web site, and check for the Nirmatrelvir file. Download the structure in the PDB format and visualize it in Chimera. Paste it here Does it look right to you? What is missing?
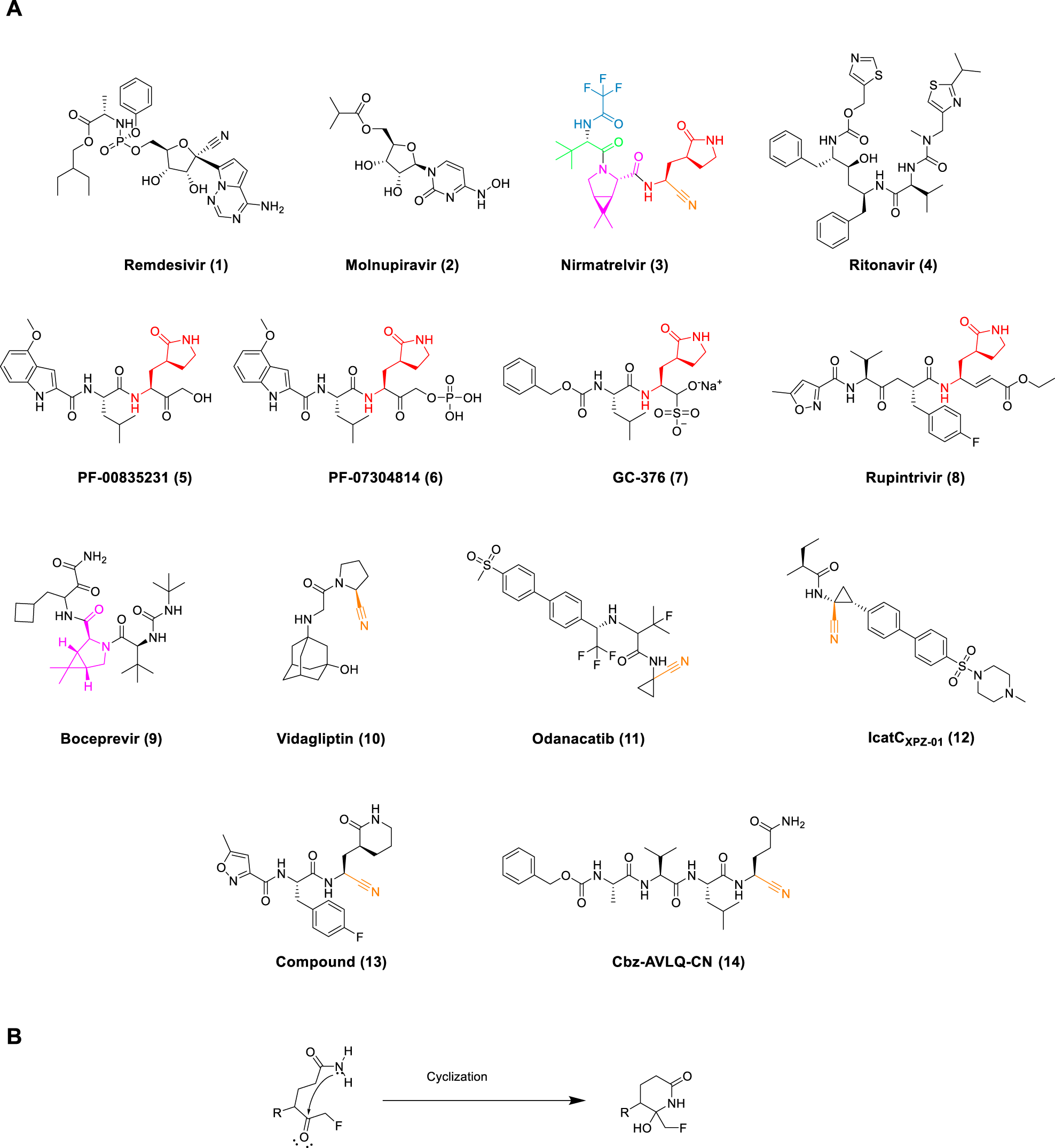 |
|---|
| Some related molecules with antiviral properties, including Nirmatrelvir (Joyce et al., 2022) |
- Try fixing the structure using openbabel. Paste the new structure as seen in Chimera.
- Can you identify the different functional groups. Discover the protein target of this molecule. Which of them is relevant for the interaction with the target?
Complex
- Can you find a structure in the PDB database that contains Nirmatrelivir with its target?
- The target is based on a conserved catalytic dyad, Can you recognize it using Chimera?
- Check the variability of the target and show in the structure where those variants at the level of aminoacids appear. Are they relevant for the function?
- What is the mode of interaction between ligand and target? Can you elaborate on why would you consider it strong and specific? How can this be related to the activity of the protein?
- Can you find information about the way Nirmatrelvir was designed? In particular, what are its precursors?
References
- You can find interesting material on COVID19 structural biology at the web of the PDB: COVID-19/SARS-CoV-2 Resources.
- Greasley, S. E.; Noell, S.; Plotnikova, O.; Ferre, R.; Liu, W.; Bolanos, B.; Fennell, K.; Nicki, J.; Craig, T.; Zhu, Y.; Stewart, A. E.; Steppan, C. M. Structural Basis for the in Vitro Efficacy of Nirmatrelvir against SARS-CoV-2 Variants. Journal of Biological Chemistry 2022, 298 (6), 101972.
- Ryan P. Joyce, Vivian W. Hu & Jun Wang The history, mechanism, and perspectives of nirmatrelvir (PF-07321332): an orally bioavailable main protease inhibitor used in combination with ritonavir to reduce COVID-19-related hospitalizations. Medicinal Chemistry Research 2022; 31, 1637-1646.
© Jordi Villà Freixa, Facultat de Ciències, Tecnologia i Enginyeries, Universitat de Vic - Universitat Central de Catalunya, 2023